
Long-range enhancer-promoter contacts work like power lines, connecting regions of the genome which are very far away from each other. Image from pixnio.com under CC PDM 1.0.
Enhancers are non-coding elements which are littered throughout the mammalian genome.
Enhancers form three-dimensional physical contacts with gene promoters, the sequence immediately upstream (before) the gene’s protein/RNA coding sequence begins.
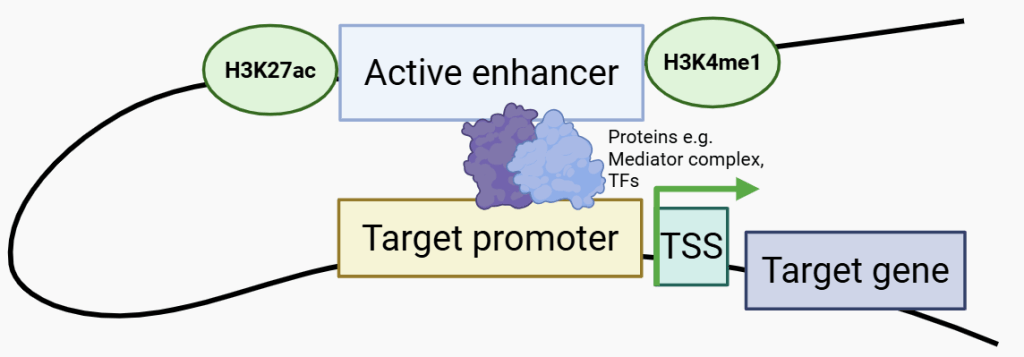
A typical (active) enhancer-promoter contact. Active enhancers are marked by permissive histone modifications such as H3K27ac and H3K4me1. Proteins, including transcription factors and the Mediator complex, allow the enhancer to contact the promoter and initiate expression of genes. TFs, transcription factors; TSS, transcription start site. Green arrow represents gene expression. Created with Biorender.com by Isabella S.
During development, thousands of enhancer-promoter contacts are established to regulate the vast network of genes required to form each part of the body. In this sense, they act like connectors between huge networks of wires which stop cables ending up in the wrong places (i.e., connecting to the wrong genes).
These contacts enable fine-tuning of gene expression, and also ensure that genes are expressed in the right time and place, which is critical during development, otherwise you’d get an arm where the head should be.
Some enhancers act over longer distances than others, with a few even reaching out to promoters on different chromosomes.
However, what makes an enhancer long- or short-acting has not yet been identified – there’s no DNA sequence or chromatin signature which defines it.
Grace Bower and colleagues publishing in Nature earlier this month have come closer to answering this critical question.

The authors specifically focused on limb development in mice embryos. Many genes governing this process are already well-characterised, one of the most famous being Sonic hedgehog (Shh), and this was indeed the authors’ primary candidate.1
The first experiment was wanted to test whether replacing the enhancer of Shh (the ZRS) with a different one impacted limb development. Deleting the ZRS has been known to abolish Shh expression, resulting in truncated limbs in mice, as early as 2005.
Indeed, they found that inserting novel enhancers into the ZRS lacked the long-range activity required to activate Shh, and consequently the limbs didn’t develop properly.
Enhancers are picky
Enhancers, like people, need a welcoming environment to be able to work productively when settling into a new role. What if the chromatin receiving the enhancers became coiled up and inactive upon insertion, preventing the new enhancer reaching all the way over to the Shh promoter?

A woman is comfortable on her sofa. When editing genomes, sequences must be inserted into accessible, “comfortable” chromatin to do their job. Created by Daniel de la Hoz, sourced from newportinstitute.com, under CC BY-NC 4.0.
The authors ruled this out. Accessibility information from ATAC-seq, which maps regions of open chromatin throughout the genome, demonstrated that the new enhancers became entrenched in open, accessible regions, suggesting that even though these enhancers were (most likely) active, they were not able to form long-range contacts.
Or maybe it was the case that the transplanted enhancers were simply incompatible with the Shh promoter? Not entirely. One enhancer, HS72, was able to activate Shh when placed immediately upstream, but not when the researchers transplanted it into the ZRS thousands of kilobases away.
So what’s going on? If amenable chromatin and a compatible promoter is not enough to make an enhancer form long-range contacts, there must be additional factors at play.
Small but powerful
Sure enough, the authors identified a small sequence adjacent to the HS72 enhancer which they suspected may support long-range activity. The sequence was found in other species too. Biologists term this evolutionary conservation, and it’s usually a good sign that the sequence serves a crucial purpose.
Remember: HS72 alone, when transplanted onto ZRS, the Shh enhancer, did not activate Shh. However, when transplanting HS72 PLUS this new sequence onto the ZRS, not only was Shh activated, but the mouse embryo developed a fully-formed limb too.
This novel sequence which converted HS72 into a long-range enhancer was termed Range Extender (REX).
The genome is not in a vacuum
Since REX had no enhancer activity itself, the authors investigated how it conferred long-range activity.
DNA sequences do not act in isolation. They need proteins, including genome architecture proteins and transcription factors, to do their job.
By scanning REX for transcription factor binding sites, they found that it contains a common binding site for LHX transcription factors, previously shown to be involved in coordinating limb growth in vertebrates.
Interestingly, the ZRS (the Shh enhancer) also had these binding sites, which, when mutated, Shh expression and limb development became stunted. Adding the REX element partially enabled the ZRS to reach its target promoter again.
See the figure below; I hope this helps. I realise there are a lot of names here, and even I got a bit confused as I was writing!
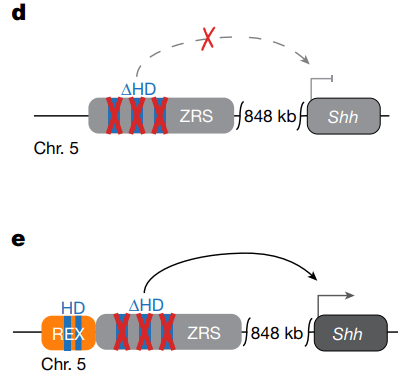
ZRS is unable to activate Shh expression with its LHX binding sites mutated (d). When the REX sequence is added, Shh expression is partially restored (e). HD, homeodomain (the sequence where the transcription factors bind); kb, kilobases. Figure from Bower et al., Nature, 2025.
This isn’t just a one-off event either. Other enhancers which work over long distances also harboured LHX binding sites, suggesting LHX may be important for governing entire networks of long-distance contacts.
“Yeah, and?”
I could talk for AGES on why enhancer-promoter contacts are relevant for development (thank you, BSc project!), but they are also relevant for our understanding of diseases, particularly cancers, in which some enhancers are “hijacked” and rewire their proper contacts, even reaching outside their defined genomic regions (topologically associated domains [TADs]) and wreaking havoc on the cell’s transcriptional landscape.2
In future, it would be interesting to see whether there are similar sequences around cancer-associated enhancers which may be involved in establishing these incorrect contacts to promoters.
Two things the authors note is that REX may not have the same function in humans, and that not all limb enhancers harbour this sequence. But these, in themselves, are other questions. Maybe we’ll find out soon.
And yes, I meant genomic distance. 😉
Notes
- But couldn’t another enhancer elsewhere in the genome take over in place of the ZRS? After all, multiple enhancers can converge on one promoter; this is called redundancy (briefly discussed in the footnotes of a previous post). The authors confirmed that, in the developing limb, only the ZRS controls Shh (see also this paper) and therefore there is no chance that some other enhancer could butt in and start regulating Shh instead, making the ZRS an ideal locus to replace to see the true effects of the new enhancer. ↩︎
- n.b.: enhancers can act outside TADs in healthy cells and still be functional, as shown recently in fruit flies. ↩︎
Discussion point
Aside from chromatin accessibility, why might an enhancer not be compatible with a certain promoter?

Leave a reply to RNA previously dismissed as “noise” now shown to produce functional proteins – The Chromatin Digest Cancel reply